Check this out: Family Tree Spreadsheets
Latest Articles and Tutorials
- How To Identify Rare Surnames In Your Family Tree
 When you’re building your family tree, the rarer surnames are often the easiest to research. My grandmother was a Smith which makes that ancestral line a challenge to explore. My early research focused on marriages in the Smith line to people with less common last names. This tutorial shows you how to use a spreadsheet … Read more
When you’re building your family tree, the rarer surnames are often the easiest to research. My grandmother was a Smith which makes that ancestral line a challenge to explore. My early research focused on marriages in the Smith line to people with less common last names. This tutorial shows you how to use a spreadsheet … Read more - 4 Ways To Get A List Of Surnames From Your Family Tree
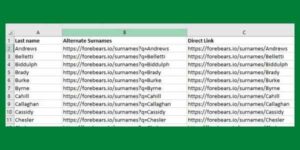 If your family tree is on Ancestry.com or another online website, there’s no simple way to export a list of surnames. This article shows you several different methods to get a full list of people from your family tree into a spreadsheet. We then show you how to turn that into a distinct list of … Read more
If your family tree is on Ancestry.com or another online website, there’s no simple way to export a list of surnames. This article shows you several different methods to get a full list of people from your family tree into a spreadsheet. We then show you how to turn that into a distinct list of … Read more - Copy-And-Paste Your Ancestry Tree To A Spreadsheet
 There are several different ways to get a list of names from your family tree. Some methods use desktop software or require a little technical knowledge. But if your tree is online on Ancestry, there is one simple way to get a list of people into a spreadsheet. It’s time-consuming with larger trees but it … Read more
There are several different ways to get a list of names from your family tree. Some methods use desktop software or require a little technical knowledge. But if your tree is online on Ancestry, there is one simple way to get a list of people into a spreadsheet. It’s time-consuming with larger trees but it … Read more - How To Split Names In Excel
 Have you got a long list of people’s names in Excel and you’re not sure how to sort or filter them by surname? The challenge occurs when the first and last name are in a single cell. You may also have middle names jammed in there. And worse, there may be a prefix (Mrs.) or … Read more
Have you got a long list of people’s names in Excel and you’re not sure how to sort or filter them by surname? The challenge occurs when the first and last name are in a single cell. You may also have middle names jammed in there. And worse, there may be a prefix (Mrs.) or … Read more - Last Names Starting With M (Over 600 Names)
 Are you looking for a comprehensive list of last names beginning with “M”? We’ve used the 2000 and 2010 U.S. Census to compile over six hundred surnames that start with the letter M and had at least five thousand people with the name In the table below, you’ll see the total number of bearers for … Read more
Are you looking for a comprehensive list of last names beginning with “M”? We’ve used the 2000 and 2010 U.S. Census to compile over six hundred surnames that start with the letter M and had at least five thousand people with the name In the table below, you’ll see the total number of bearers for … Read more - Create A Four-Generation Family Tree In Google Sheets
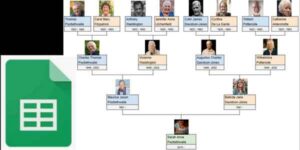 This tutorial takes you step-by-step through creating a four-generation family tree in Google Sheets. A four-generation tree goes back to eight great-grandparents. This tutorial builds a tree that prints on a single landscape page. If you’re too busy for the twelve steps in this tutorial, jump down to the end to grab our “done for you” Google Sheets template bundle. … Read more
This tutorial takes you step-by-step through creating a four-generation family tree in Google Sheets. A four-generation tree goes back to eight great-grandparents. This tutorial builds a tree that prints on a single landscape page. If you’re too busy for the twelve steps in this tutorial, jump down to the end to grab our “done for you” Google Sheets template bundle. … Read more - Make a Word Cloud From Your Family Tree
 A word cloud is a visual representation of text. The size of each word in the cloud is proportional to its frequency. When we use word clouds to represent our family tree, we are usually making one from either: In this tutorial, I show two ways to make a word cloud based on last names … Read more
A word cloud is a visual representation of text. The size of each word in the cloud is proportional to its frequency. When we use word clouds to represent our family tree, we are usually making one from either: In this tutorial, I show two ways to make a word cloud based on last names … Read more - How To Convert A GEDCOM File To Spreadsheet (Script Provided)
 Do you want to convert a GEDCOM file into a spreadsheet with every person nicely formatted in rows? This tutorial walks through how to do this for free. You’ll be working with a script or piece of code, but don’t worry if you’re not familiar with programming. I’ll provide the script in a way that … Read more
Do you want to convert a GEDCOM file into a spreadsheet with every person nicely formatted in rows? This tutorial walks through how to do this for free. You’ll be working with a script or piece of code, but don’t worry if you’re not familiar with programming. I’ll provide the script in a way that … Read more - GEDCOM – An Introduction For Beginners
 This article explains the basics of the GEDCOM file format that is used copy family trees between different platforms and software. I answer all the basic questions, with a gentle introduction to some of the more complex topics. What Is GEDCOM And How Does It Work? Suppose that you download your family tree from Ancestry.com … Read more
This article explains the basics of the GEDCOM file format that is used copy family trees between different platforms and software. I answer all the basic questions, with a gentle introduction to some of the more complex topics. What Is GEDCOM And How Does It Work? Suppose that you download your family tree from Ancestry.com … Read more - Create A Six Generation Family Tree In Google Sheets
 A six-generation tree goes up to 32 great-great-great-grandparents. Our layout in Google Sheets prints on a single landscape page. That’s perfect for bringing to libraries and archives. If you’re too busy for the twelve steps in this tutorial, jump down to the end to grab our “done for you” Google Sheets template bundle. What The 6-Generation Pedigree Tree Looks Like … Read more
A six-generation tree goes up to 32 great-great-great-grandparents. Our layout in Google Sheets prints on a single landscape page. That’s perfect for bringing to libraries and archives. If you’re too busy for the twelve steps in this tutorial, jump down to the end to grab our “done for you” Google Sheets template bundle. What The 6-Generation Pedigree Tree Looks Like … Read more








